colorbar and axis labels overlayed - how to fix? (or "how to give parameters to save that i can give to show?")
This helped me to fix the label on my colorbar, and I have consistent fonts now, too. yay.
however two issues remain:
# Region where contour lines should be plotted
def region_to_include(x,y):
return (x-.5)^2+(y-.5)^2>0.01 and (x+.5)^2+(y-.5)^2>0.01 and (x-.5)^2+(y+.5)^2>0.01 and (x+.5)^2+(y+.5)^2>0.01
# Region to be excluded
def region_to_exclude(x,y):
return not region_to_include(x,y)
#=================== visual plausibility check
pic=phi_ges.subs({
Q : 1e-9,
d_e : 1,
z_s : .25,
z_c : .5,
epsilon_m : 76,
epsilon_s : 21.75,
epsilon_0 : 8.854187817e-12
})
from matplotlib import rc, rcParams
rc('text',usetex=True)
rc('font',**{'family':'serif','serif':['Computer Modern']})
c = contour_plot(pic, (x,-1,1), (z,-1,1), region=region_to_include, contours= 30, plot_points=200, cmap=colormaps.jet, colorbar=True, axes_labels=['$x$ in [m]','$z$ in [m]'], typeset='latex', frame=True, axes=False)
plt=c[0]
opt = plt.options()
opt['colorbar_options']['label'] = 'Potential in $V$'
plt.set_options(opt)
c[0] = plt
# Plot of the region to be excluded
r = region_plot(region_to_exclude, (x,-1,1), (z,-1,1),
plot_points=300, incol="gray", bordercol="black", borderwidth=1)
# Final plot
show(c + r,)
bild=c + r
bild.save('potentialfeld.pdf')
gives me a plot like this:
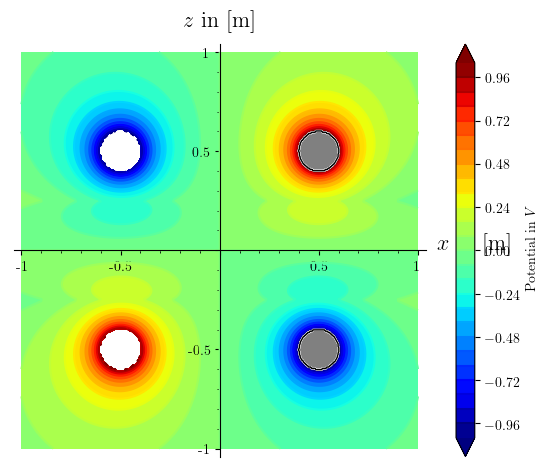
why does my frame=True, axes=False get ignored in the first plot command? It does work if i add those parameters to a separate show command, like show(c+r, frame=True, axes=False), but then i can't save it like that.
saving the plot manually from within the viewer that show() spawns gives me this pic, as i want it. but i would like to be able to save it from sage directly.
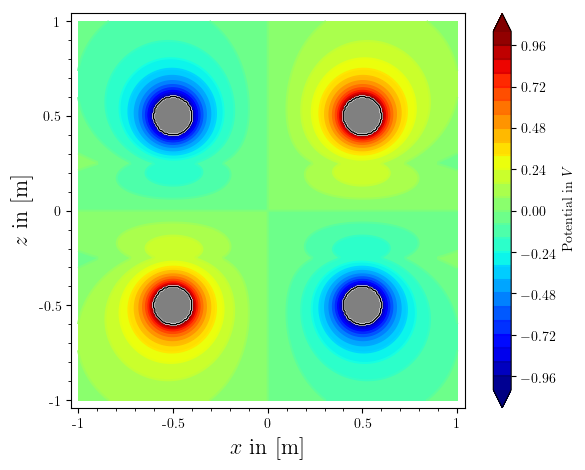
how to fix?
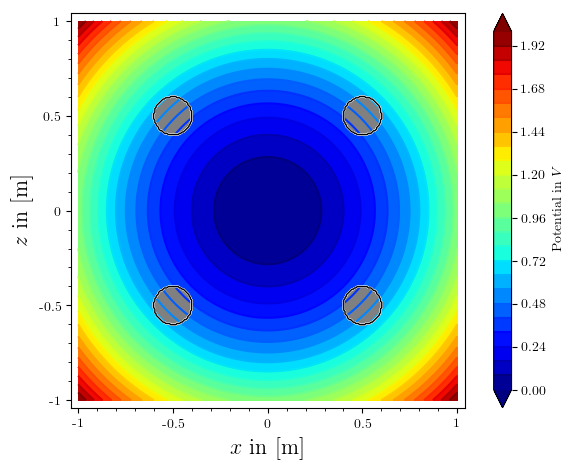

before i had also asked about an issue with the poles on the left being not greyed out, i found and fixed the typo.