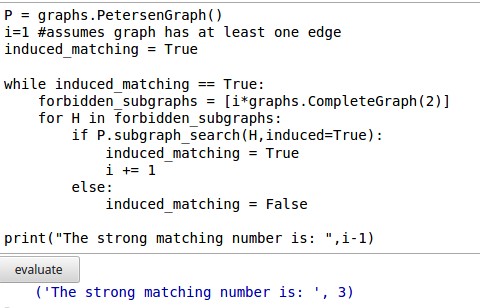
One solution is to use Knuth's dancing links algorithm which is implemented in Sage. Assuming the vertices of the graph are integers in the range(number_of_vertices), the method below will work:
from sage.combinat.matrices.dancing_links import dlx_solver
def find_matching(G):
rows = [[a,b] for (a,b,_) in G.edges()]
dlx = dlx_solver(rows)
solution = dlx.one_solution(ncpus=1)
if solution:
return [rows[a] for a in solution]
else:
return None
Then, on the Petersen graph, it gives:
sage: G = graphs.PetersenGraph()
sage: matching = find_matching(G)
sage: matching
[[2, 7], [0, 1], [5, 8], [3, 4], [6, 9]]
sage: G.plot(edge_colors={'red':matching})
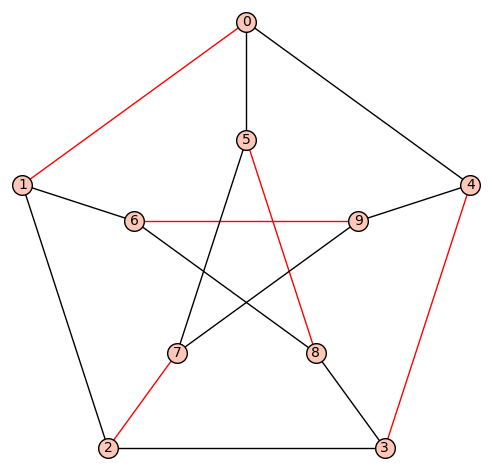
Another solution is to reduce the problem to SAT and use Sage SAT solvers or you may also review the ticket I wrote recently about the reduction of dancing links instance to SAT instance.